Enzymes, as you have seen in another post, are biocatalysts. They help speed up biological reactions. While you just saw the basics, let us explore enzymes at a deeper level.
Properties of Enzymes
| Specificity A particular enzyme will only bind to one specific substrate. This property is exploited in drug discovery. | Temperature Sensitivity Enzymes are proteins, and they degrade when exposed to really high temperatures. Each enzyme has a specific temperature at which it is most active |
| Cannot function independently They require coenzymes or cofactors to function effectively. examples include Mg2+. | Most enzymes are proteins, with a few exceptions While most enzymes are proteins, there are a few exceptions- Ribozymes, which are RNA. they are basically RNAs with catalytic activity and can be evident in many biological reactions. |
Enzyme Classification
| Type | Function/ Mechanism of action | Examples |
|---|---|---|
| Oxidoreductases | carry out redox reactions | oxidase, dehydrogenase |
| Transferases | involved in group transfers | kinase |
| Hydrolases | cleavage of chemical bonds through hydrolysis | nuclease, protease |
| Lyases | non hydrolytic cleavage of chemical bonds | decarboxylase, aldolase |
| Isomerases | converts one isomeric form to the other | triose phosphate isomerase |
| Ligases | joining two substrate molecules through covalent bonds | DNA ligase |
Enzyme activity
- As you have learnt in my previous post, enzymes catalyze biological reactions by reducing the Gibbs free energy of activation. this makes it easier for the reaction to reach it’s transition state faster
- Michaelis- Menten equation gives the rate (v) at which these enzyme catalyzed reactions take place
- When you plot substrate concentration (on X axis) against reaction time (on Y axis), you get a hyperbolic curve
- You can see this hyperbolic curve as two parts-
- the linear portion- indicating rapid product formation
- the steady portion- where the enzyme uses up the substrate or the enzyme loses it’s activity
Enzyme kinetics
- This is usually studied using a widely accepted Michaels-Menten model/ equation
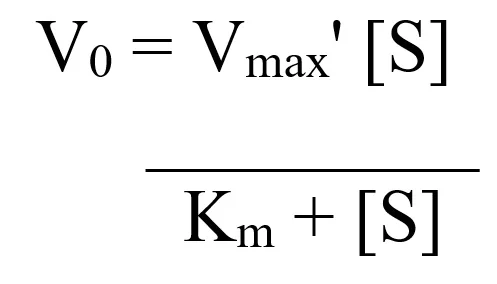
- Km is the michaels-mentens constant, it is the measure of affinity of a substrate for the enzyme
Kinetic parameters
- This can give us an idea of the maximum rate which an enzyme can attain and how easily it gets saturated with a substrate
- An enzyme’s kinetic parameters can tell us how an enzyme will behave in a cell and how it will respond to changes
- The michaels-mentens constant Km is yet another key parameter that tells us the substrate concentration required to reach half it’s Vmax
Lineweaver Burkplot
- Also known as double reciprocal plot
- It transforms the Michael mentens equation into a straight line in order to calculate Km and Vmax easier
- This graphical method plots reciprocal of reaction rate (1/V) against reciprocal of substrate concentration (1/[S])
- Through this, we can identify the Vmax (the maximum rate when enzyme is saturated with substrate) and Km (the substrate concentration at which the reaction rate is half of Vmax
Factors affecting enzyme activity
A lot of factors could affect enzyme activity. these include-
- temperature- they influence the rate at which the enzyme catalyzes the reaction. A reaction requires collision of 2 molecules- an enzyme and a substrate. at low temperatures, collision is less frequent than at higher temperatures. However, if the temperature exceeds a certain limit,, the enzyme will degrade because they are proteins. An optimal temperature at which human enzymes function is 37 degrees C.
- pH- just like temperature, enzymes need an optimal pH . Any deviation from this will affect its functioning as they lose their 3D structure and change in shape of their active sites. So, the substrate cannot bind to the active site and the enzyme is inefficient. each enzyme has it’s own optimal pH depending on its location in our body.
- substrate concentration- rate of enzyme catalyzed reaction initially increases with increase in substrate concentration. However, at a point when all enzymes are saturated with substrate, rate of enzyme catalyzed reaction will plateau.
- enzyme concentration- More enzyme molecules mean more active sites for substrate binding, which leads to an increase in reaction rate. However, if the amount of substrate is limited, increasing enzyme concentration beyond a certain point will have no effect on enzyme activity, since there will be a surplus of unnecessary enzyme molecules.
- Presence of inhibitors- they can come in three different ways
- competitive inhibition- inhibitor is very similar to the substrate it is competing with – with regard to shape and size. If this binds to the enzyme the substrate will not be able to bind to your enzyme and the desired product will not be formed.
- non- competitive inhibition- inhibitor binds to a site other than the active site called the allosteric site. this changes the shape of the enzyme and affects it’s activity regardless of the activity.
- feedback inhibition- when enough product is formed, it inhibits the enzyme activity in order to prevent excess product formation.
Multisubstrate reactions
- Involves one enzyme, many substrates
- They involve complex rate equations to know how the substrates bind and in what order
- These types of reactions are physiologically relavant as many biological reactions involve multiple substrates
- These reactions take place through two mechanisms-
- sequential reaction- all the susbtrates must bind to the enzyme before any product is released. they can be of two types
- ordered sequential- substrates bind in a specific sequence
- unordered sequential- substrates bind in a random order
- Ping-pong mechanism- one substrate binds to enzyme— product is released—– next substrate binds—- next product released
- sequential reaction- all the susbtrates must bind to the enzyme before any product is released. they can be of two types
- Kinetics of such reactions become slightly more complicated
- We obtain initial velocity by keeping one of the substrate concentration constant and varying the other
- their Lineweaver Burk plots have distinct patterns to identify if they work through sequential or ping pong mechanism- sequential if the lines are intersecting and ping pong if the lines are parallel
Enzyme Units
- It is the unit of enzyme catalytic activity- International Unit (IU)
- It refers to the amount of enzyme that catalyzes the conversion of 1umole of substrate per minute under specific conditions
Catalytic strategies
- Covalent catalysis- the catalyst forms a temporary bond with the substrate
- serine residue and its substrate involves covalent bond formation
- Acid-Base catalysis- involves proton transfer to or from a substrate
- amino acid side chains in enzymes act as acid acceptors or donor
- Metal-ion catalysis- where a metal ion stabilizes the charges or mediates redox reactions
- A metal ion, when bound to the substrate can
- Proximity catalysis- where the catalyst brings substrates together in the correct orientation
- two molecules which are meant to be together or join with each other but they cant as they are just floating in the air. Enzymes brings them both and connects them together
- Electrostatic catalysis- the electrical charges of amino acids in the enzyme can help stabilize the transition state
Mechanism of RNase
- As the name suggests, it is a small enzyme (144 amino acids) which degrades RNA (and not DNA) by cleaving the phosphodiester bonds in them
- They may be endoribonucleases (cuts single or double stranded RNA) or exoribonucleases (cut the terminal ends of an RNA)
- Most important residues in the enzyme- 2 Prolines, 2 Histidines (His12, His119) and Lysine (Lys 41)
- Substrate for this enzyme- 2′,3′-cyclic phosphodiester on 3′ side of a pyrimidine residue
- Mechanism: acid-base catalysis
- Base- His12. it gets a proton from oxygen of the substrate, thereby facilitating its attack on a phosphorus
- This attack displaces a nucleoside
- Acid- His119. it donates a proton to 5′ oxygen
- This displaces a nucleoside
- Both the displaced products are released into the solvent
- Lys41 along with phenylalanine will stabilize this transition state
- It is the most studied enzyme due to its physiological function- it is a digestive enzyme secreted by pancreas
Mechanism of Chymotrypsin
- It is a serine protease which cleaves peptide bonds- includes serine as a nucleophile in the enzyme site
- Physiological significance- its a component of the pancreatic juice
- Involves a catalytic triad where it cleaves- tyrosine, tryptophan, phenylalanine
- Substrate- aromatic or bulky non polar side chains
- Catalytic triad- His57, Ser195, and Asp102
- Reaction mechanism- ping pong mechanism
- Enzyme forms a covalent bond with the substrate- displaces a stable moiety
- Enzyme-substrate complex, also known as intermediate is formed
- Intermediate reacts with water, displaces remaining part of the substrate and reforms original enzyme
- In this whole reaction,
- Serine provides nucleophile (O atom)
- His acts as a catalyst and activates serine
- Asp stabilizes the protonated Histidine
Mechanism of Carbonic Anhydrase
- It is a hydrolytic enzyme- catalyzes the addition or removal of water to a substrate
- It catalyzes the reversible conversion of carbon dioxide to bicarbonate (aka carbonic acid)
- It is physiologically important as it is involved in acid-base balance and CO2 transport
- Substrate- CO2 molecule
- Active site- Zinc-bound hydroxide (ZnOH)
- The ZnOH attacks CO2 displacing a zinc-bound bicarbonate ion
- A water molecule enters the enzyme site and removes the bicarbonate ion from zinc
- Rate limiting step – proton from a zinc bound water is transferred to an active site residue-His64
- This step will regenrate the ZnOH, release the proton, and forms bicarbonate
Mechanism of Lysozyme
- It is an antibacterial enzyme
- Active site Glu35 and Asp52
- Substrate- Peptidoglycan
- It hydrolyzes beta 1,4-glycosidic bonds between N-acetyl muramic acid (NAM) and N-acetyl glucosamine (NAG)
- NAM and NAG are present in peptidoglycan layers of gram positive bacteria
- This weakens the cell wall leading to cell lysis and eventually bacterial cell death due to osmotic pressure
- Mechanism
- Glu35- acid catalyst- donates proton to oxygen of glycosidic bond
- Asp52- nucleophile- forms a temporary covalent bond with the carbon atom of NAM
- At this stage, water enters the active site— Glu35 abstracts proton from water making oxygen in water nucleophilic
- Oxygen attacks carbon that Asp52 is bound to and breaks the bond between the C and Asp52
- Now that Glu35 is protonated and Asp52 is deprotonated, and the two resulting products are released
Regulation of Enzymes
Enzymes cannot and need not be produced at the same level everytime. Their production needs to be controlled. Under certain circumstances, they need to be induced (produce more enzyme) and in some they need to be repressed (produce less enzymes). There are different mechanisms to do this-
- Allosteric regulation- effector molecules bind to an allosteric site (different from the enzyme’s active site) thereby changing the conformation of the enzyme either inducing it or repress it
- Covalent modification- chemical groups like phosphate are added to or removed from certain amino acids in the enzyme altering it’s activity
- Regulatory inhibitors- they are effector molecule which bind to the enzyme and reduce or block its activity. They can either directly bind to the active site competing with the enzyme (competitive inhibition) or bind to an allosteric site and prevent the substrate from binding to the enzyme (non-competitive inhibition)
- Feedback inhibition- This happens in a metabolic pathway. Once enough end product is formed, it binds to to the enzyme and represses it preventing further product formation.


Thanks for sharing!